Previous Projects in Detail
Assessment of psychiatric disorders using pH-sensitive imaging methods: Casey Johnson, PhD, Postdoctoral Fellow
Casey Johnson, PhD

A growing area of interest in psychiatric research is the role of pH in various disorders, for example bipolar and panic. Baseline pH as well as pH dynamics may be fundamental to these disorders. To better understand these mechanisms, methods that can probe brain pH noninvasively are needed. A number of MR methods have been shown to be pH sensitive, including 31P and 1H (lactate) spectroscopy, T1rho imaging, and chemical-exchange saturation transfer (CEST). Spectroscopy techniques are the most common approach to noninvasively assess brain pH. However, these methods have coarse spatial resolution and require long scan times. The methods for pH-sensitive T1rho and CEST imaging are fairly new and have not been extensively demonstrated in vivo. However, these methods have the potential to enable imaging of the entire brain with high spatial and temporal resolution within a reasonable scan time. My project involves the use of these pH-sensitive methods, as well as the development and assessment of improved techniques, to image patients with psychiatric disorders, specifically bipolar and panic. The primary goal of this work is to identify pH differences in these patients versus normal controls. A secondary goal is to develop pH-sensitive MR imaging methods that are practical for human imaging at 3T.
T1rho Precision Calculator
BrainCSI: Jeff R. Yager, PhD Student
Jeff R. Yager

Magnetic Resonance Spectroscopy is a technique used in brain imaging to analyze the brain’s chemical content. Spectroscopy data is typically collected with a single voxel (SV) method or a multiple voxel method called Chemical Shift Imaging (CSI). A software tool, BrainCSI, is being developed to help process data obtained in these experiments. The metabolite concentrations are most commonly reported as ratios of two metabolites, but the BrainCSI tool attempts to report absolute concentrations. One current limitation absolute concentrations in spectroscopy is the large voxel size ( ~ 1 cm3). The large voxel size causes multiple tissues types to be contained inside a voxel. Brain metabolites concentrations are known to vary according to the tissue type, most notably that cerebral spinal fluid (CSF) is void of most metabolites of interest. Therefore, it is important to determine each tissue partial volume of the spectroscopy voxel. The BrainCSI tool classifies the anatomical image into different tissue classes and determines the partial volume. The point spread function of the spectroscopy voxel in CSI is also significant and is corrected for in BrainCSI. Finally, in 2D CSI and SVS, there is a chemical shift that is metabolite dependant that needs to be corrected.
Brain pH Changes: Hye Young Heo, PhD Student and Wen Li, PhD
Hye Young Heo

Brain pH has been shown to fluctuate as a result of brain activity. It has been difficult to measure such changes in vivo due to lack of methods for non-invasively measuring pH with high spatial and temporal resolution. In this study we performed a series of experiments to access the ability and sensitivity of T1 relaxation in the rotating frame (T1ρ) to measure pH changes in the mouse and human brain evoked by systemically manipulating carbon dioxide or bicarbonate. Furthermore, T1ρ detected a localized acidosis in the human visual cortex induced by a flashing checkerboard. This was confirmed with 31P spectroscopic measurements. In addition, 1H spectroscopy showed a significant elevation of the lactate signal providing a mechanism for the acidosis. These studies will open significant opportunities for understanding the role of pH-dependent signaling in the brain, for understanding the importance of pH in neurological disease, and for novel diagnostic and therapeutic approaches.
Wen Li, PhD

I recently graduated from the MRI Research Lab with a PhD in Biomedical Engineering. My research was focused primarily on applying a combination of medical image processing tools into a pipeline, which provides an automated labeling to the surface of the human cerebral cortex.
The cerebral cortex is a highly convolved thin layer neural tissue located at the outermost layer of the human brain. It can be divided into anatomically and functionally meaningful sub-regions. We developed a fast and automated parcellation pipeline, which takes T1 and T2 images of the subject and automatically parcellates the cortical surface of each hemisphere into 24 well defined regions. This tool can be potentially used in studying the brain morphology related with psychiatric and neurological disorders. It will help us understand the underlying pathology that can occur when the brain gets sick. Our parcellation pipeline has been tested to be comparable with manual parcellations. It is a flexible pipeline, which can incorporate various sets of labeling systems and is faster than FreeSurfer (it takes approximately ¼ of time to get similar results).
BRAINS3 Software Development: Steve Dunner, Programmer
Steve Dunner

My primary role at MRRF is working on the creation of the BRAINS3 software package, a joint venture with software engineers in the Dept. of Psychiatry. This is a suite of tools to facilitate systematic analysis of MRI datasets. This analysis is performed both dynamically by neuroscientists as well as automatically by the system itself. My work focuses on the latter. I have created several tools for use in this automated processing, including, but not limited to: dynamic generation of Talairach atlases, skull-stripping/brain-masking and surface generation. My current project is a tissue classifier that will allow a user to provide an list of associated pixel value ranges and tissue types, which will then be used to accurately label the entire brain. This can be fine-tuned on a variety of parameters (primarily statistical thresholds). In my spare time I like to volunteer at the hospital and the Knights of Columbus, study topics in computer science and engineering and avoid cleaning my apartment at all costs.
Automated Building Block Assignments for Hexahedral Finite Element Meshing: Austin J. Ramme, MD/PhD Student
Austin J. Ramme, MD/PhD

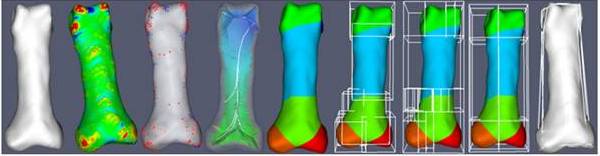
Finite element analysis has allowed for investigations of the human musculoskeletal system and orthopaedic implant design. IA-FEMesh, an open-source hexahedral meshing software package, provides tools to develop volumetric meshes on a patient-specific basis. Part of the IA-FEMesh meshing protocol relies on manual placement of building blocks around a virtual model. As the complexity of the surface increases (e.g. phalanx versus cervical vertebrae), the number of required blocks increases and hence the time required to create the block structure increases. The Automated Building Block Algorithm incorporates principles from differential geometry, combinatorics, statistical analysis, and computer science to automatically generate a building block structure for a given surface without prior information. The finalized building block structure is imported into IA-FEMesh to generate a hexahedral mesh for finite element analysis. Two metrics were used to judge the quality of the resulting finite element mesh: minimum element Jacobian and minimum element volume. As a final check of mesh quality, simulated material properties, boundary conditions, and initial conditions were applied to the mesh, and the mesh was analyzed using the ABAQUS® finite element solver.
It is common to perform group analysis of diffusion tensor data based on rotationally invariant scalar indices like fractional anisotropy (FA), relative anisotropy (RA) etc. All of the subjects for the study are mapped to a common coordinate system, often defined by an atlas image. The transformation between the space of the acquired diffusion weighted images and the atlas space is defined via an image registration procedure. The resulting transformation is then applied to the scalar images where the voxel values within the atlas space are interpolated using conventional techniques such as linear interpolation. To perform group analysis of diffusion tensors, it is necessary to interpolate the tensor representation as well as rotate the diffusion tensors to keep the tensors consistent with the tissue reorientation. Higher order diffusion models like the Q-Ball model (QBI) or high order tensor model that are gathered from high angular resolution diffusion imaging (HARDI) data, also need to be reoriented during spatial normalization. This research is focused on developing a generalized me
The steps of the Automated Building Block Algorithm are illustrated above: (i) reorientation of the bone surface model, (ii) surface feature identification, (iii) point selection filter, (iv) medial axis analysis, (v) surface segmentation, (vi) raw building block definitions, (vii) combinatorial building block reorganization, (viii) building block manipulation, and (ix) finalized building block vertex projection.
Spatial Normalization of Diffusion Models and Tensor Analysis: Madhura A. Ingalhalikar, PhD, Biomedical Engineering
Madhura A. Ingalhalikar, PhD
It is common to perform group analysis of diffusion tensor data based on rotationally invariant scalar indices like fractional anisotropy (FA), relative anisotropy (RA) etc. All of the subjects for the study are mapped to a common coordinate system, often defined by an atlas image. The transformation between the space of the acquired diffusion weighted images and the atlas space is defined via an image registration procedure. The resulting transformation is then applied to the scalar images where the voxel values within the atlas space are interpolated using conventional techniques such as linear interpolation. To perform group analysis of diffusion tensors, it is necessary to interpolate the tensor representation as well as rotate the diffusion tensors to keep the tensors consistent with the tissue reorientation. Higher order diffusion models like the Q-Ball model (QBI) or high order tensor model that are gathered from high angular resolution diffusion imaging (HARDI) data, also need to be reoriented during spatial normalization. This research is focused on developing a generalized method for spatial normalization of diffusion models. Following below are the aims of this project:
- Develop a generalized tool for spatial normalization of tensors and other higher order diffusion models.
- Implement non-linear intersubject registration using a multichannel registration scheme.
- Validate the above methods primarily on tensors and then on higher order diffusion models.
- Develop tensor analysis tools.

Expectation Maximization Based Bone Segmentation: Austin J. Ramme, MD/PhD Student
Medical imaging technologies have allowed for in vivo exploration and evaluation of the human musculoskeletal system. Three-dimensional bone models generated using image segmentation techniques provide a means to optimize individualized orthopaedic surgical procedures using engineering analyses. Traditionally, manual raters have performed the segmentation of bone from medical images. However these techniques are not clinically practical due to the extensive time and human intervention that is required. This study aims to develop an automated preprocessing and segmentation method to accurately, efficiently, and reliably separate bony regions of interest. Others in our laboratory are working towards producing automatic hexahedral meshes from the surface definitions appropriate for surface contact analysis using the finite element method. Our ultimate goal is to automate the preprocessing, segmentation, and mesh generation procedures to provide a clinically useful means of surgical analysis and planning.
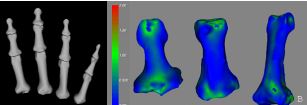
Figure (A): Surface models of the anterior aspects of twelve phalanx bones generated using the Expectation Maximization algorithm for image segmentation. Figure (B): Euclidean distance map comparison between physical bone surface scans and the Expectation Maximization algorithm based segmentations of the index finger.
Imaging of the Lung: Bill Kearney, PhD
Bill Kearney, PhD

We are developing pulse sequences and methodologies to image lung structure and function. The structure and function of the lung has been investigated using magnetic resonance with hyperpolarized He and Xe. Due to the high cost and complexity of using these agents, we are seeking more versatile and economical alternatives. Fluorine imaging is a promising approach that may replace exotic nuclei in both structural and functional pulmonary studies. Fluorine has a high intrinsic MR sensitivity, and a number of biologically benign reagents are available to investigate properties of both the gas and tissue volumes. Fluorine imaging may also provide information about the gas/tissue interface that is not accessible by any other method.
Correlated Spectroscopy and Quantitative Susceptibility Mapping: Cameron Cushing, PhD
Cameron Cushing, PhD

Despite advances in treatment capabilities, biological understanding, and imaging technologies, a diagnosis of glioblastoma remains a death sentence for patients. Pharmacological ascorbate therapy has promise as a safe adjuvant to standard of care treatment for primary glioblastoma as demonstrated by multiple clinical trials. Pharmacological ascorbate is hypothesized to act as a pro-drug for the production of hydrogen peroxide and superoxide via an iron-catalyzed auto-oxidation reaction. Auto-oxidation of ascorbate reduces Iron(III) to Iron(II). Methods that quantitate ascorbate in vivo or monitor the reduction of the labile iron pool by ascorbate may provide important information for managing ascorbate therapy. Direct quantitation of endogenous ascorbate has previously been demonstrated in vivo via magnetic resonance spectroscopy. Correlated spectroscopy (COSY), a two-dimensional spectroscopic method, has shown promise in separating and quantitating multiple chemical species in vivo. Preliminary data suggest that ascorbate may be quantitated through this method as well. Iron can be measured by the MR imaging techniques T2* relaxometry and quantitative susceptibility mapping (QSM). Iron measurement via MR imaging has historically neglected to examine the influence of redox active metal pools on these imaging methods. My preliminary results suggest that reduction of Iron(III) may alter these images in vivo.
Thoracic Imaging: Stanley Kruger, PhD
Stanley Kruger, PhD

I am interested in MRI as a tool for thoracic imaging applications generally. Specifically, I have two primary areas of thoracic interest. The first is investigating lung structure and function in order to better assess regional ventilation and perfusion, especially as it relates to disease and therapy. In pursuit of this aim, I am interested in the use of hyperpolarized gas contrast agents (He-3 or Xe-129), Flourine Imaging, and Oxygen-Enhanced Proton MRI. My second area of interest is to collaborate on development of an MR-only radiation oncology suite for tumor treatment in the chest. Ultimately, I am interested in overcoming the challenges of respiratory motion, as well as the removal of the need for multi-modal acquisitions in treatment planning.
MR Imaging using Deep Learning: Hemant Kumar Aggarwal, PhD
Hemant Kumar Aggarwal, PhD

Hemant is developing algorithms for fast MR imaging using deep learning techniques. During his Ph.D., he has worked on compressed sensing, dictionary learning, low-rank matrix recovery, and joint sparse recovery related techniques. His first project was on compressive multispectral imaging that is an extension of the color image demosaicing problem often encountered in the design of low-cost single-sensor cameras. He had developed a uniform multispectral filter array to capture multiple bands using single-sensor and a demosaicing algorithm to reconstruct the full multispectral image from the undersampled raw image. He also worked on spatio-spectral total-variation based method for hyperspectral image denoising problem in the mixed noise scenario where mixed noise may have Gaussian noise as well as sparse noise that includes line strips, shot noise, and impulse noise. He further considered the blind source separation problem also known as the unmixing problem that focuses on identifying constituent classes and their fractions present at each pixel in a hyperspectral image.
Accelerating dynamic free-breathing cardiac MRI: Sunrita Poddar, PhD candidate
Sunrita Poddar, PhD

I work on the acquisition and reconstruction of ungated dynamic cardiac data in the free-breathing mode. We have acquired these cardiac images in a time comparable to a breath-held scan, and found that the reconstruction quality is comparable to that obtained from a breath-held acquisition. In addition, our proposed technique is feasible in critically ill patients and also paediatric patients who are unable to hold their breath or follow breath-holding instructions. A short summary of this work can be found here and a more detailed description can be found in our paper. This work has been conducted in the context of CINE imaging. Our future work involves extending our developed algorithms to enable ungated free-breathing perfusion imaging and parameter mapping. Our vision is to develop a comprehensive cardiac exam which can conduct ungated free-breathing CINE, perfusion imaging and parameter mapping in a single short scan.
Magnetic Resonance Imaging (MRI) reconstruction design for fast Imaging acquisition: Yassir AlBaqqal, PhD student
Yassir AlBaqqal, PhD

The ultimate goal of my research is to develop novel algorithms to reconstruct heavily undersampled sparse imaging. The designed schemes aim to achieve a higher patio-temporal resolution, signal-to-noise ratio and coverage in multidimensional multichannel MRI. In addition to improving patients comfort and compliance while imaging under the MRI device, the new developed schemes will allow patients with arrhythmia problems, pediatrics and obese people to breath freely without the need for any breath-hold scans. Shortening examination periods also reduces patient's stress, lowers the entire visit to the clinic and finally lowers the associated economic costs. Rapid imaging acquisitions will also allow for efficient extraction of quantitative information needed for the patients diagnosis eg. tumor characterization and veins blockages through myocardial perfusion MRI. Current application of interests include real time CINE MRI and contrast changing perfusion MRI.
Structured signal reconstruction from incomplete data: Sampurna Biswas, PhD candidate
Sampurna Biswas, PhD

The main thrust of my research is on the development of algorithms for the reconstruction of structured signals from incomplete data. These algorithms are expected to have applications in medical imaging and sonar array processing. The underlying theme has been designing sampling schemes to observe the signal, devising transformation of the signal based on its inherent structure and then deriving theoretical recovery guarantees for the chosen sampling scheme and recovery algorithm. Signals collected from a patient undergoing a MRI scan, are correlated and share same sparsity patterns, across time. The image reconstruction can be posed as a low rank and sparse signal recovery. There exists sampling scheme in the literature that solves this recovery in two steps of subspace estimation. The guarantee for this scheme to provide an exact solution was not available. I proposed a generalized two-step low rank and joint sparse signal recovery with theoretical guarantees (in terms of number of measurements required) for each step. This is impactful in designing the acquisition scheme for a two-step recovery in dynamic MRI applications like CINE, myocardial perfusion and brain parametric mapping MRI.
T1ρ Imaging: Nana Owusu, PhD Student
Nana Owusu, PhD

My work entails a relaxation parameter (contrast) known as T1rho or longitudinal relaxation in the rotating frame. T1rho is a form of contrast in magnetic resonance imaging and this contrast is due to the exchange of protons in biological fluids. There is no consensus as to which species of protons in biological fluids contributes greatly to T1rho signal (e.g. due to pH, glucose, other macromolecules). Understanding more about the signal contribution may help clinicians in assessing treatment or explain more about ailments linked to abnormal metabolism like Huntington's disease, bipolar disorder I, cancer and epilepsy. I am also working on GE pulse sequence development for T1rho imaging.















